For the second week of our Women in Science month, we are pleased to present Dr. Kerstin Howe. She is a German computational biologist working on genomic sequences (1).
Biography
Kerstin Howe was born in Germany. She studied Biology at the Ruhr University of Bochum in Germany where she had her diploma in 1994 and after a year working in industry returned to university to obtain a PhD in genetics in 1999 (1,2). After she finished her PhD, she worked for one year at this university in the general and molecular botany group on assembling and annotating mitochondrial sequences.

Kerstin always preferred computational biology to lab work and counted herself lucky to join the Wellcome Sanger Institute in Cambridge, UK, as a Senior Computer Biologist in August 2000, initially annotating the human genome (2). She soon moved on to become a programmer in the wormbase team, annotating and building the reference databases for C. elegans genome (1–3).
When the Sanger Institute started a new flagship project sequencing the zebrafish genome in 2001, Kerstin got tasked with its gene annotation and from 2002 led the Zebrafish Genome Analysis Team.
This work led to Kerstin’s involvement in founding of the Genome Reference Consortium maintaining the human and mouse reference genome assemblies, in 2008, together with relevant teams at the NCBI, Washington University St. Louis (now the McDonnell Genome Institute) and EMBL-EBI (4–6).
Kerstin is still involved in the GRC and since 2021 heads the genome assembly production of the Tree of Life Department at the Sanger Institute, generating highest quality reference assemblies for multiple large biodiversity genomics projects. The largest of these is the Darwin Tree of Life project, aiming at sequencing and annotating all 70,000 eukaryotic species in the British Isles (4–6).
Her contribution to zebrafish research
In 2001, the Wellcome Sanger Institute started the Zebrafish Genome Project and Kerstin Howe was immediately a member of the project (7).
It soon turned out that assembling the zebrafish genome was much more complicated than initially anticipated and therefore in 2002 a team was founded to address these issues, with Kerstin taking the lead.
During her time with the Zebrafish Genome Project at the Sanger Institute, Kerstin and her team developed new methods of assessing and correcting assembly data, whilst releasing continually improved zebrafish assemblies.
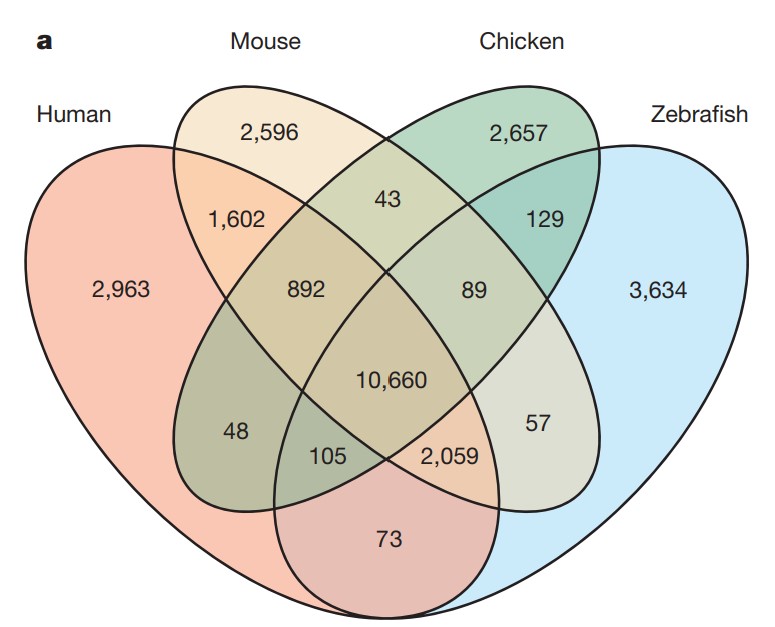
With the 9th iteration of the zebrafish assembly, Zv9, in 2013, the assembly was deemed mature enough to be published. This article, “The zebrafish reference genome sequence and its relationship to the human genome” in 2013 was of relevant importance to understand gene homologies between zebrafish and humans (10).
This research performed the assembly of the zebrafish genome and provided evidence of more than 26 ‘000 protein-coding genes (10). The comparison with the human genome shows approximately 70% of human genes having at least one zebrafish homology.
The zebrafish genome assembly has since been fully adopted by the GRC and Kerstin’s team have continued to improve the genome assembly, with the current latest iteration being GRCz11.
Conclusion
Through genome assembly generation and genome analyses, Kerstin Howe participated in making it possible to use zebrafish as a model organism for human development and disease.
By being at the Head of Production Genomics in the Tree of Life Programme, Kerstin contributes to the generation of assemblies for the EarthBiogenome Project, biology’s moonshot to sequence, catalog, and characterize the genomes of all of Earth’s eukaryotic biodiversity.
Dr. Kerstin Howe contributes also in the conservation of endangered species. By reconstructing the genome of endangered species, she has highlighted the importance of genetic diversity on the survival of a species. The more genetic diversity a species has, the better it is able to adapt to change and survive (5).
References
- Howe, Kerstin – Wellcome Sanger Institute [Internet]. [cited 2023 Mar 6]. Available from: https://www.sanger.ac.uk/person/howe-kerstin/
- Kerstin Howe (0000-0003-2237-513X) [Internet]. ORCID. [cited 2023 Mar 7]. Available from: https://orcid.org/0000-0003-2237-513X
- Kerstin Howe | LinkedIn [Internet]. [cited 2023 Mar 7]. Available from: https://www.linkedin.com/in/kerstin-howe-36736b25/
- Dashboard – Darwin Tree of Life [Internet]. [cited 2023 Mar 6]. Available from: https://schools.darwintreeoflife.org/profile/
- Completing the puzzle of life on Earth [Internet]. [cited 2023 Mar 8]. Available from: http://innovationstories.sanger.ac.uk/completing-the-puzzle-of-life-on-earth/
- Darwin Tree of Life – Reading the genomes of all life: a new platform for understanding our biodiversity [Internet]. [cited 2023 Mar 7]. Available from: https://www.darwintreeoflife.org/
- ZFIN Person: Howe (fka Jekosch), Kerstin [Internet]. [cited 2023 Mar 6]. Available from: https://zfin.org/ZDB-PERS-001130-2
- Zebrafish Genome Project – Wellcome Sanger Institute [Internet]. [cited 2023 Mar 7]. Available from: https://www.sanger.ac.uk/data/zebrafish-genome-project/
- ZFIN Lab: Sanger Centre Zebrafish Project [Internet]. [cited 2023 Mar 7]. Available from: https://zfin.org/action/profile/view/ZDB-LAB-001206-2
- Howe K, Clark MD, Torroja CF, Torrance J, Berthelot C, Muffato M, et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature. 2013 Apr 25;496(7446):498–503.
First image: Completing the puzzle of life on Earth [Internet]. [cited 2023 Mar 8]. Available from: http://innovationstories.sanger.ac.uk/completing-the-puzzle-of-life-on-earth/
Read and approved by Kerstin Howe: Kerstin Howe | LinkedIn https://www.linkedin.com/in/kerstin-howe-36736b25/
