The shift to 3D cell culture, with spheroids, organoids, and assembloids, is opening new doors in biological research, offering models that better mimic human tissues and disease. Yet as these models become more intricate, so do the challenges: ensuring uniformity, establishing reproducible and scalable production, etc. This month, we explore how automation is helping researchers meet these challenges head-on, making advanced 3D culture more accessible and reliable than ever.
Introduction
The quest for more predictive and physiologically relevant in vitro models has propelled 3D cell culture, encompassing spheroids, organoids, and assembloids, to the forefront of biological research and drug discovery. As New Approach Methodologies (NAMs) gain traction, these advanced 3D models are bridging the gap between oversimplified 2D cultures and complex in vivo systems, offering unprecedented insights into human tissue function and disease mechanisms.
Yet, as the complexity of these models grows, so do the challenges. Manual handling of 3D cultures introduces variability, limits throughput and reproducibility, which are critical hurdles for researchers aiming to translate discoveries into reliable, scalable solutions. How can we ensure uniformity across hundreds of samples? What does it take to standardize the production for high-throughput screening? And how do we ensure physiological relevance?
Automation is emerging as the key solution, transforming these challenges into opportunities. By integrating cutting-edge robotic systems and workflows, researchers are unlocking new levels of precision, reproducibility, and scalability in 3D cell culture. This month, we explore how automation is reshaping the landscape of advanced 3D models, highlighting innovations like Bionomous’ collaboration with Celvivo, which exemplifies the power of technology to propel biological research forward.
Defining the models
The shift from 2D to 3D cell culture has unlocked new possibilities in biological research, with spheroids, organoids, and assembloids offering increasingly sophisticated models of human tissues. Each of these models brings unique strengths and challenges to the table, depending on the research goals.
- Spheroids are the simplest form of 3D culture: spherical aggregates of cells formed through spontaneous adhesion, often derived from cell lines, primary cells, or tumor tissues. Their rapid formation (typically within 2–3 days) and cost-effectiveness make them a staple for high-throughput applications, such as drug screening, toxicity testing, and tumor microenvironment studies (1, 2). However, their simplicity comes with limitations. Spheroids lack the structural complexity of native tissues, often consisting of just a few cell layers (proliferating cells on the outside, quiescent cells in the middle, and a hypoxic or necrotic core) which can restrict their ability to fully mimic physiological conditions (2).
- Organoids: represent a significant leap forward. These self-organizing 3D structures, derived from stem cells or tissue, recapitulate the architecture and function of specific organs (3, 4, 5). Cultured using methods like submersion (matrix embedding vs suspension), air-liquid interfaces, bioreactors or microfluidics, organoids are invaluable for disease modeling, regenerative medicine, and personalized drug development. Yet, their complexity introduces challenges, including the need for precise culture conditions, difficulties in standardization, and limitations in vascularization and long-term maintenance.
- Assembloids: push the boundaries even further by combining multiple organoids or cell types to model interactions between tissues. These advanced systems are particularly useful for studying neural migration, circuit formation, and multi-organ crosstalk, offering a modular approach to reconstructing complex biological processes (6, 7).
The following table summarizes the key differences between spheroids and organoids, highlighting their unique features and applications:
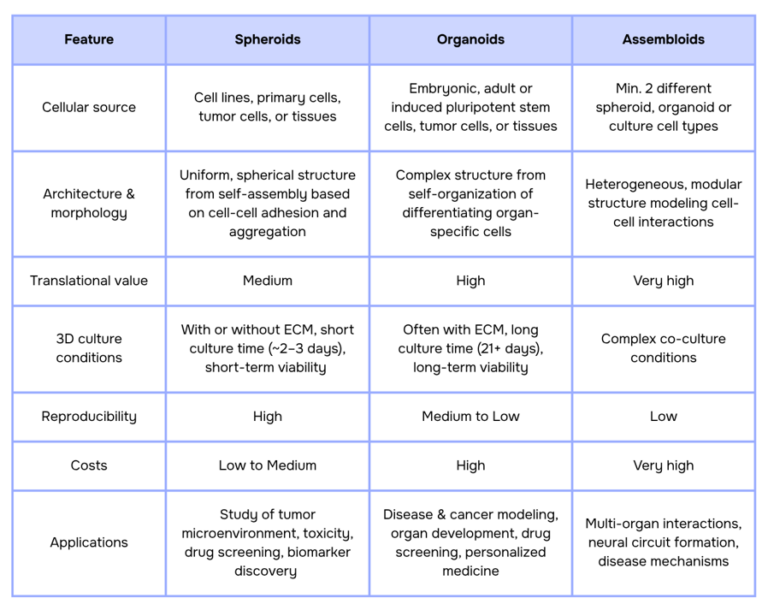
While spheroids, organoids, and assembloids offer powerful tools for biological research, their complexity introduces key challenges. Issues such as oxygen and nutrient diffusion, morphology consistency, size heterogeneity, and scalability can hinder reproducibility and limit their potential. In the next sections, we will explore these current challenges and how innovative solutions are addressing them.
Current challenges
Despite their transformative potential, 3D culture systems still face several fundamental limitations that restrict their reproducibility, scalability, and broader adoption. These challenges are not isolated but deeply interconnected, arising from both biological complexity and technical constraints. Understanding and addressing these limitations is essential to unlock the full potential of spheroids, organoids, and related models.
Oxygen & Nutrient diffusion
The absence of a vascular system in most 3D cultures creates a critical bottleneck. Oxygen and nutrients can only diffuse ~100–200 µm from the surface, leading to hypoxic cores and necrosis in larger structures (8). This limitation not only distorts experimental outcomes, such as drug efficacy or metabolic studies, but also restricts long-term cultures, which are essential for modeling chronic diseases or developmental processes. While microfluidic systems can partially address these issues by mimicking vascular flow, their global adoption remains limited by technical and cost barriers.
Standardization & Reproducibility
Achieving consistency in 3D cultures remains a major challenge, as size, morphology, and internal structure vary widely depending on the cell type and application. For example, tumor-derived spheroids and patient-derived organoids (PDOs) are often deliberately size-controlled within ~40–100 µm for high-throughput or microfluidic applications to ensure uniform diffusion and assay compatibility (9). In contrast, more mature or structurally organized organoids typically exceed 150–200 µm, with size often used as a proxy for structural organization (10). Similarly, liver organoids used for toxicity testing are commonly maintained within ~100–500 µm to preserve viability and function (11), while brain organoids can grow from ~400 µm to several millimeters, reflecting their increasing architectural complexity (12). This variability highlights that organoid size is not a fixed parameter, but one that is inherently linked to both biological origin and experimental design.
This heterogeneity directly impacts diffusion gradients, drug penetration, and cellular behavior, making standardization both difficult and highly application-dependent. Without uniform protocols for generation, culture, and analysis, reproducibility suffers. Even minor variations in seeding density, extracellular matrix composition, or culture conditions can lead to significant differences in organoid size, shape, and internal organization. As a result, comparing data across experiments remains challenging, limiting the broader adoption of 3D models in high-throughput applications such as drug screening and personalized medicine.
Scalability
Manual protocols for generating and maintaining 3D cultures remain a major bottleneck for their broader adoption. These workflows are inherently time-consuming, labor-intensive, and highly operator-dependent, making them prone to variability and human error (13). As demand increases for high-throughput applications such as drug screening and personalized medicine, scaling up production while maintaining consistency in organoid size, morphology, and function becomes particularly challenging. Organoid workflows often involve multiple complex and manual steps, from cell seeding and matrix embedding to media exchange and monitoring, each introducing potential variability.
Beyond culture generation, downstream handling, including transfer, positioning, and analysis, also remains largely manual and difficult to standardize. This limits throughput and introduces batch-to-batch variability that is difficult to control or trace, complicating cross-experimental comparisons and large-scale studies. As a result, the lack of scalable and standardized workflows continues to hinder reproducibility and restrict the integration of 3D models into routine and industrial applications.
The critical role of automation
Building on these challenges, recent work shows that improving organoid reproducibility and scalability requires a broader shift toward engineering-controlled systems, where variability is addressed across materials, spatial organization, and process workflows and not only through biological optimization (14). While advances in defined matrices and spatial patterning help reduce intrinsic variability, scaling these systems to high-throughput applications introduces new operational bottlenecks.
In this context, automation plays a central role by enabling standardization, process control, and integration across the organoid workflow, from production to analysis.
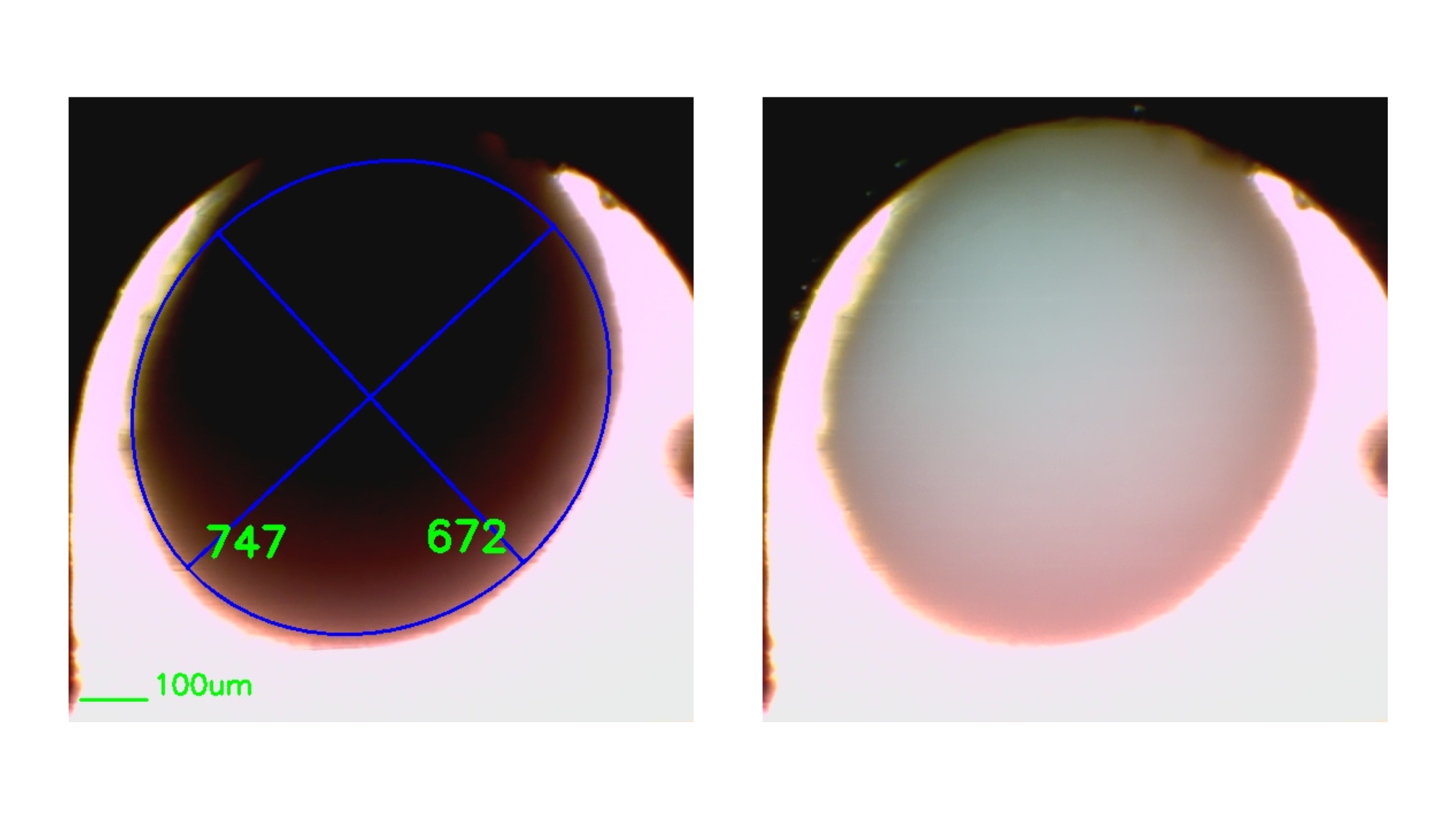
- Quality control (QC): Automated imaging and analysis enable objective, quantitative assessment of organoid size, morphology, and development. Coupled with machine learning, these systems allow real-time, label-free evaluation and standardized selection criteria.
- Reproducibility through process control: Automation reduces operator-induced variability by standardizing key steps such as cell seeding, media exchange, and environmental regulation, which is critical for systems highly sensitive to small fluctuations in culture conditions.
- Throughput and scalability: As scalable production platforms (e.g., microwell arrays, microfluidic systems, bioreactors) generate increasing numbers of organoids, automation ensures that downstream operations like feeding, monitoring, and analysis can match this throughput.
- Automated handling and integration: Beyond culture, automated handling technologies enable precise manipulation, sorting, and positioning of organoids, reducing labor and enabling end-to-end workflow integration.
- Operational efficiency: Automation reduces hands-on time, minimizes human error, and improves consistency, making large-scale organoid applications more feasible in both research and industrial settings.
A representative example is the integration of dynamic culture systems with automated selection technologies. In workflows combining CelVivo bioreactors with Bionomous platforms, organoids can be cultured under controlled conditions and then automatically sorted based on size and morphology, enabling the selection of homogeneous populations for downstream applications.
Ultimately, automation is not an isolated solution but a key enabling layer within a broader engineering framework, supporting the transition from variable, small-scale organoid cultures to reproducible, scalable, and application-ready systems.
Conclusions & Outlooks
The choice between spheroids, organoids, and assembloids ultimately depends on the application, as each model offers different levels of complexity, physiological relevance, and scalability. However, across all systems, key challenges remain—particularly variability, limited standardization, and constraints in scaling.
Addressing these limitations requires a shift toward engineering-driven approaches, combining defined materials, controlled microenvironments, and scalable production platforms. Within this framework, automation plays a central role by enabling standardized workflows, integrated quality control, and high-throughput processing.
Looking ahead, the convergence of biomaterials, microfabrication, and automation is set to transform 3D cultures into reproducible and scalable systems. The development of integrated, end-to-end workflows, from production to analysis, will be essential to move from experimental models to reliable, application-ready platforms, ultimately enabling broader adoption in drug discovery, precision medicine, and regenerative medicine.
References
- Živković, Z., & Opačak-Bernardi, T. (2025). An Overview on Spheroid and Organoid Models in Applied Studies. Sci, 7(1), 27. https://doi.org/10.3390/sci7010027
- Logos Bio. (2026, March 11). 3D cell culture: Differences between spheroids and organoids and their analytical strategies. https://logosbio.com/3d-cell-culture-differences-between-spheroids-and-organoids-and-their-analytical-strategies/
- Meudec, L., Goudarzi, N., Mariette, X., & Nocturne, G. (2025). Lessons from organoid engineering for rheumatic disease. Joint Bone Spine, 92(3), 105604. https://doi.org/10.1016/j.jbspin.2025.105948
- Gunti, S., Hoke, A. T. K., Vu, K. P., & London, N. R., Jr (2021). Organoid and Spheroid Tumor Models: Techniques and Applications. Cancers, 13(4), 874. https://doi.org/10.3390/cancers13040874
- Jones, J. (2023, April 21). Spheroids vs. organoids: Which 3D cell culture model is best for you? Novus Biologicals. https://www.novusbio.com/antibody-news/spheroids-vs-organoids-which-3d-cell-culture-model-is-best-for-you
- Onesto, M. M., Kim, J. I., & Pasca, S. P. (2024). Assembloid models of cell-cell interaction to study tissue and disease biology. Cell stem cell, 31(11), 1563–1573. https://doi.org/10.1016/j.stem.2024.09.017
- Park, Y., et al. (2021). Three-dimensional, multifunctional neural interfaces for cortical spheroids and engineered assembloids. Science Advances, 7(22), eabf9153. https://doi.org/10.1126/sciadv.abf9153
- Pham, M. T., Pollock, K. M., Rose, M. D., Cary, W. A., Stewart, H. R., Zhou, P., Nolta, J. A., & Waldau, B. (2018). Generation of human vascularized brain organoids. Neuroreport, 29(7), 588–593. https://doi.org/10.1097/WNR.0000000000001014
- Nitschke, C., Phan, C., Souto, Y., et al. (2025). Patient-derived tumor organoids highlight the potential of precision medicine in managing pancreatic ductal adenocarcinoma. International Journal of Cancer, 157(4), 760–772. https://doi.org/10.1002/ijc.35443
- Surina, Tanggis, Suzuki, T., Hisata, S., Fujita, K., Fujiwara, S., Liu, F., Fukushima, N., Suzuki, T., Mato, N., & Hagiwara, K. (2023). Patient-derived spheroids and patient-derived organoids simulate evolutions of lung cancer. Heliyon, 9(3), e13829. https://doi.org/10.1016/j.heliyon.2023.e13829
- Moon, H. R., Mun, S. J., Kim, T. H., Kim, H., Kang, D., Kim, S., Shin, J. H., Choi, D., Ahn, S. J., & Son, M. J. (2024). Guidelines for Manufacturing and Application of Organoids: Liver. International journal of stem cells, 17(2), 120–129. https://doi.org/10.15283/ijsc24044
- Ozaki, T., Ohta, N., Ma, J., & Hirano, M. (2025). 360° size-adjustable microelectrode array system for electrophysiological monitoring of cerebral organoids. Frontiers in bioengineering and biotechnology, 13, 1596009. https://doi.org/10.3389/fbioe.2025.1596009
- Lampart, F. L., Iber, D., & Doumpas, N. (2023). Organoids in high-throughput and high-content screenings. Frontiers in Chemical Engineering, 5, 1120348. https://doi.org/10.3389/fceng.2023.1120348
- Kim, D., Youn, J., Kim, J., et al. (2026). From organoid culture to manufacturing: Technologies for reproducible and scalable organoid production. npj Biomedical Innovations, 3, 12. https://doi.org/10.1038/s44385-025-00054-6
